MDR 101 - Part 5 - Interpretation
MDR is a data mining approach that uses constructive induction or attribute construction to facilitate the detection of interactions in the absence of main effects. A common criticism of data mining and machine learning methods is that they take inputs and produce an output with little understanding of the 'black box' in the middle that processes the information provided. The black box phenomenon can make models discovered by data mining methods difficult to interpret. We have placed a priority recently on providing tools that provide a statistical interpretation of MDR models that we hope will improve our ability to develop a biological interpretation.
The first thing I do is to study the graphical model of the best MDR model. If the model only involves two attributes the interpretation is fairly simple. The first question to ask is whether the distribution of high-risk (dark-shaded) and low-risk (light-shaded) genotype combinations looks nonlinear. That is, do they vary within and between rows and columns? A strong main effect will show up as a column and/or row that is all high-risk or all low-risk. In other words, the effect of the genotype doesn't vary across the other genotypes. This of course becomes more difficult in the higher dimensions which is why we now rely on information theory approaches.
An important concept in data mining is information gain which is based on measures of entropy. That is, how much information is gained about case-control status from knowledge about genotypes at one or more SNPs? Stated another way, how much entropy in case-control status is removed by considering genotype? For the purposes of an MDR analysis we would like to know how much information about case-control status is gained by combining two or more SNPs using the MDR attribute construction function. More specifically, do we gain information above and beyond that provided by each SNP individually? This is what Jakulin and others call interaction information. We have taken these ideas and methods and implemented the interaction dendrogram in the MDR software for interpreting MDR models. For more details please see our paper in the Journal of Theoretical Biology from 2006. Additional information about interaction information can be found in our 2011 Genetic Epidemiology paper and several in press papers that extend these methods to 3-way interactions (will post these soon).
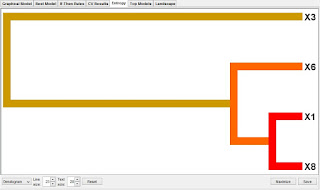
The idea is simple. If combining two or more SNPs using MDR gives a positive information gain then there is evidence for a synergistic interaction. If the combination of SNPs gives a negative information gain then information is lost which happens when there is redundancy or correlation (e.g. linkage disequilibrium). If there is no gain or loss then you can conclude the SNPs have independent effects. The dendrogram (default) in the Entropy tab is constructed in the following way. First we compute the information gain for each SNP in the summary table and then each pairwise MDR combination. These information gain values are then inverted and a distance matrix constructed such that pairs of SNPs with stronger interactions have a smaller distance. This distance matrix is then used to carry out a hierarchical cluster analysis resulting in an interaction dendrogram. The shorter the line connecting two attributes the stronger the interaction. The color of the line indicates the type of interaction. Red and orange suggest there is a synergistic relationship (i.e. epistasis). Yellow suggests independence. Green and blue suggest redundancy or correlation. Thus, you can very quickly scan the dendrogram to identify the epistasis effects in your MDR analysis. Also try the interaction graphs. These are better than the dendrograms in that they also show the marginal effects. A limitation of this analysis is that it only considers pairs of attributes. We will implement higher-order entropy analyses in a future version of the MDR software. I highly recommend that you include a dendrogram along with your best model in your presentations and publications.
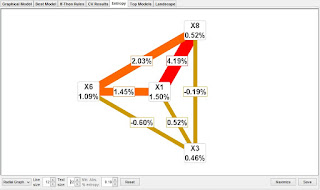
Note that a useful way to present the information analysis is in an interaction graph. These can be selected from the lower left of the Entropy tab window. Graphs have the advantage of visualizing the main effects and the interactions simultaneously. The graphs also show all the connections instead of the best that is shown in the dendrogram. We are releasing soon an MDR version 3.0 that will include additional network analysis tools.
Biological interpretation of these models will most likely be more difficult than the computation that went into the analysis. See our paper in Bioessays from 2005 for a discussion about biological inference. For biological interpretation I recommend using bioinformatics tools such as STRING for protein-protein interactions and data integration tools such as IMP that infer biological relationships across hundreds or thousands of experimental data sets from microarray studies, for example.
This post was last updated on January 20, 2013.


2 Comments:
I like it a lot! Very nicely done. Easy to find helpful information. All the best!
- compgen.blogspot.com w
spaghetti alla carbonara
Wonderful site. Love the software, but still learning how to interpret the data. The more on analysis and interpretation the better. Thank you!
Post a Comment
<< Home