An Open-Ended Computational Evolution System for the Genetic Analysis of Epistasis
Our paper on the development and evaluation of a prototype computational evolution system for epistasis analysis has been accepted for oral presentation at the EvoBIO'08 conference in Naples, Italy in March of 2008. The peer-reviewed paper will be published by Springer in Lecture Notes in Computer Science. Email me after Jan. 1st for a preprint. A complete list of accepted papers for the conference can be found here. This paper was inspired by the Banzhaf et al. review that distinguishes 'artificial evolution' and computational evolution' and fits with our theme of using expert knowledge to guide stochastic search algorithms for genetic analysis. I have included Figure 1 below the abstract.
Moore, J.H., Andrews, P.C., Barney, N., White, B.C. Development and Evaluation of an Open-Ended Computational Evolution System for the Genetic Analysis of Susceptibility to Common Human Diseases. Lecture Notes in Computer Science, in press (2008).
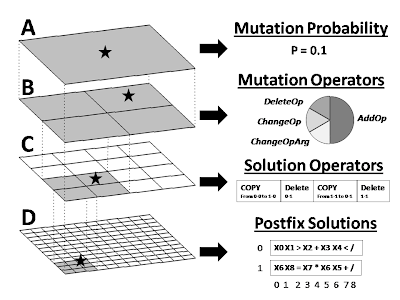
Figure 1. Visual overview of our prototype computational evolution system for discovering symbolic discriminant functions that differentiate disease subject from healthy subjects using information about single nucleotide polymorphisms (SNPs). The hierarchical structure is shown on the left while some specific examples at each level are shown on the right. The top two levels of the hierarchy (A and B) exist to generate variability in the operators that modify the solutions. Shown in C is an example set of operators that will perform recombination on the two solutions shown in D. As illustrated in B, there is a 0.50 probability that a mutation to the recombination operator in C will add an operator thus making this particular operator more complex. This system allows operators of any arbitrary complexity to modify solutions. Note that we used a 24x24 grid of solutions in the present study. A 12x12 grid is shown as an illustrative example.